For three individuals with clonal expansions that can be estimated using our methods, we have longitudinal data to orthogonally validate these estimates, which is included here. Additionally, for 13 clones with a driver gene matching a driver gene in the single cell data, but without a match to a specific clone, we include this longitudinal data as well.
Format
A data.frame containing all the information needed
- Sample.ID
The individual's ID
- Age
Individual's age at the various sampling times
- VAF
The variant allele frequency at the various sampling times for the clone of interest
- Gene
Gene or genes with mutation that identifies the clone
- Protein
Protein affected by the mutation
- cellType
The type of cells used for sequencing
- cloneName
The name we use for the clone to match to single cell data, if applicable.
References
These datasets were generated and annotated in: Williams et al. 2022 Fabre et al. 2022
Examples
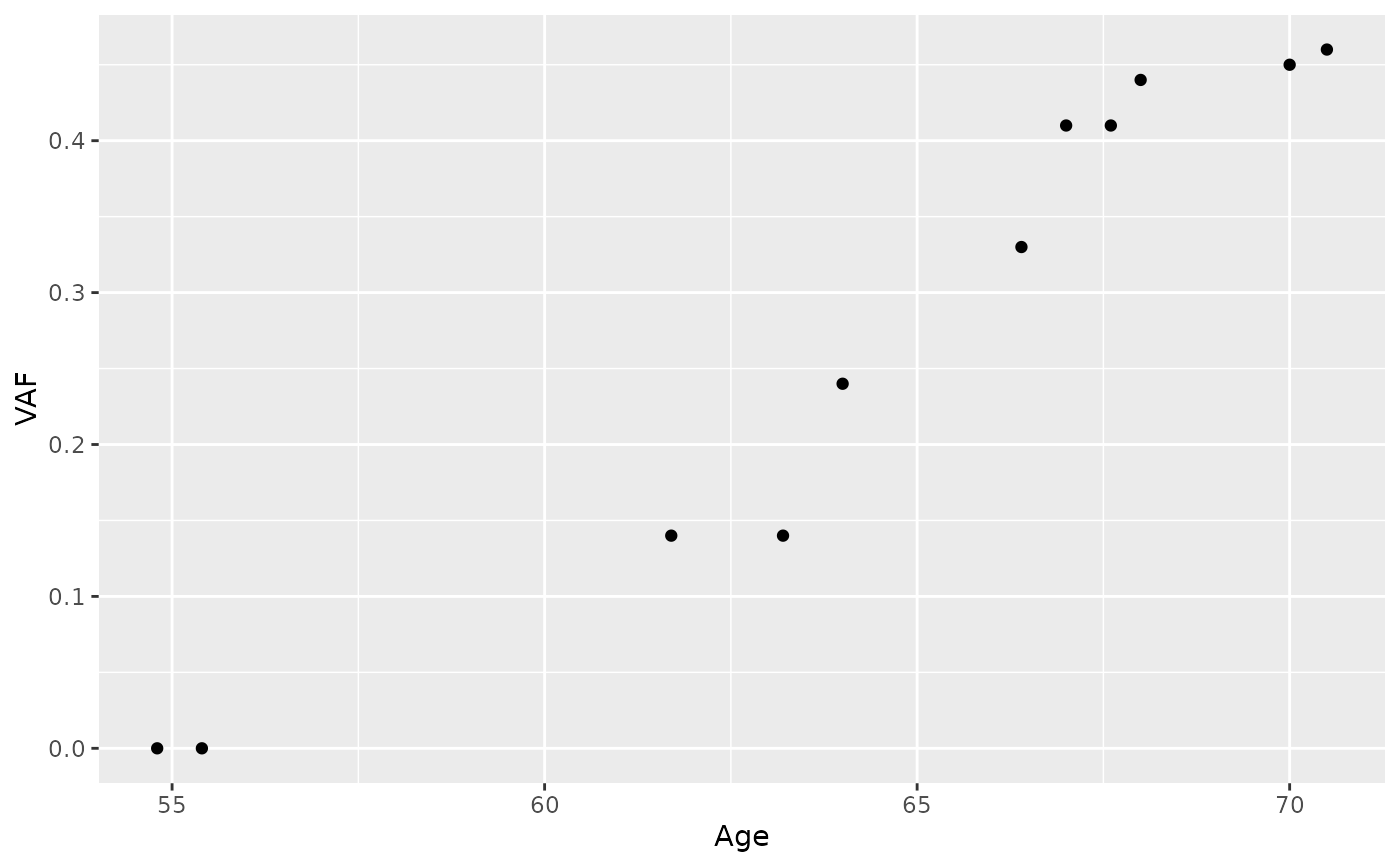
# Plot longitudinal data from PD9478
library(ggplot2)
ggplot(longitudinalData[longitudinalData$Sample.ID == "PD9478", ]) +
geom_point(aes(x = Age, y = VAF))